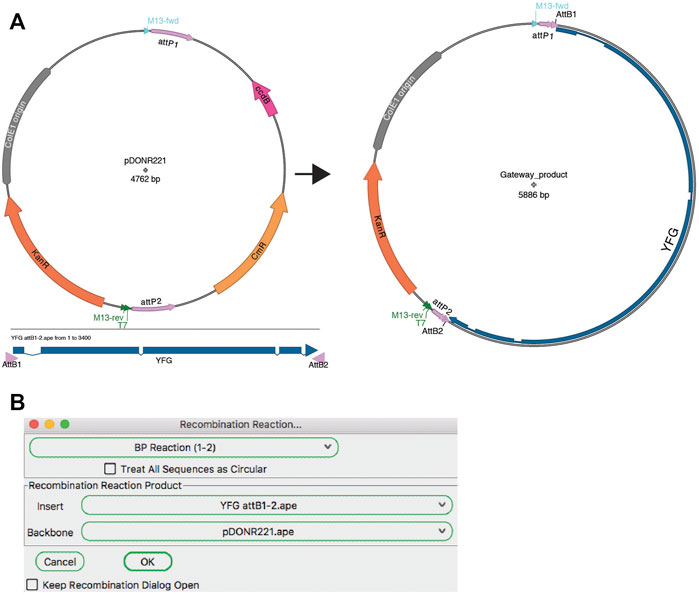
Research at Ambion has found that typically more than half of randomly designed siRNAs provide at least a 50% reduction in target mRNA levels and approximately 1 of 4 siRNAs provide a 75-95% reduction. If desired, you may modify this target site selection strategy to design siRNAs with other dinucleotide overhangs, but it is recommended that you avoid G residues in the overhang because of the potential for the siRNA to be cleaved by RNase at single-stranded G residues. In Elbashir's and subsequent publications, siRNAs with other 3' terminal dinucleotide overhangs have been shown to effectively induce RNAi. This is also compatible with using RNA pol III to transcribe hairpin siRNAs because RNA pol III terminates transcription at 4-6 nucleotide poly(T) tracts creating RNA molecules with a short poly(U) tail. 
(1) that siRNAs with 3' overhanging UU dinucleotides are the most effective. 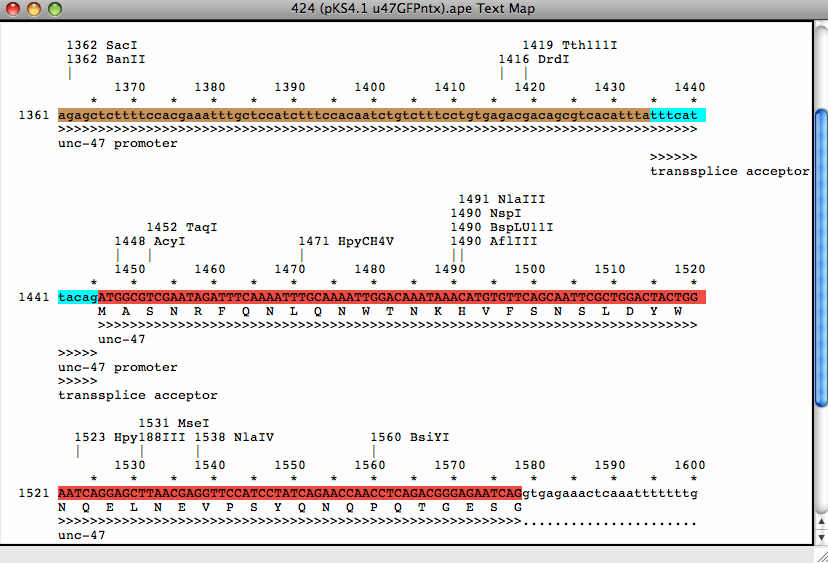
This strategy for choosing siRNA target sites is based on the observation by Elbashir et al. Record each AA and the 3' adjacent 19 nucleotides as potential siRNA target sites. Find 21 nt sequences in the target mRNA that begin with an AA dinucleotide.īeginning with the AUG start codon of your transcript, scan for AA dinucleotide sequences. Corresponding siRNAs can then be chemically synthesized, created by in vitro transcription, or expressed from a vector or PCR product.ġ. If you prefer to design your own siRNAs, you can choose siRNA target sites in a variety of different organisms based on the following guidelines.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
